NPs Basic Information

|
Name |
(6aR,10Z,11aS,11bR)-10-(1-hydroxyethylidene)-7,7-dimethyl-6a,7,11a,11b-tetrahydro-6H-pyrrolo[1',2':2,3]isoindolo[4,5,6-cd]indole-9,11(2H,10H)-dione
|
| Molecular Formula | C20H20N2O3 | |
| IUPAC Name* |
(2R,3S,5Z,9R)-5-(1-hydroxyethylidene)-8,8-dimethyl-7,16-diazapentacyclo[9.6.1.02,9.03,7.015,18]octadeca-1(17),11(18),12,14-tetraene-4,6-dione
|
|
| SMILES |
C/C(=C/1\C(=O)[C@@H]2[C@@H]3[C@@H](CC4=C5C3=CNC5=CC=C4)C(N2C1=O)(C)C)/O
|
|
| InChI |
InChI=1S/C20H20N2O3/c1-9(23)14-18(24)17-16-11-8-21-13-6-4-5-10(15(11)13)7-12(16)20(2,3)22(17)19(14)25/h4-6,8,12,16-17,21,23H,7H2,1-3H3/b14-9-/t12-,16+,17+/m1/s1
|
|
| InChIKey |
CNZIQHGDUXRUJS-PTNHGACKSA-N
|
|
| Synonyms |
alpha-Cyclopiazonic acid; CHEBI:17734; CHEBI:22450; CHEMBL468766; 83136-88-3; (6aR,10Z,11aS,11bR)-10-(1-hydroxyethylidene)-7,7-dimethyl-6a,7,11a,11b-tetrahydro-6H-pyrrolo[1',2':2,3]isoindolo[4,5,6-cd]indole-9,11(2H,10H)-dione; MFCD00167445; Cyclopiazonic acid from Penicillium cyclopium; Cyclopiazonic acid, 98%; CHEMBL480627; HMS1791B05; HMS1989B05; HMS3402B05; BDBM50453613; BDBM50527068; CCG-208182; DB07604; NCGC00163130-01; NCGC00163130-02; NCGC00163130-03; (6aR,11aS,11bR)-10-acetyl-11-hydroxy-7,7-dimethyl-2,6,6a,7,11a,11b-hexahydro-9H-pyrrolo[1',2':2,3]isoindolo[4,5,6-cd]indol-9-one; alpha-Cyclopiazonic acid 100 microg/mL in Acetonitrile; 6H-Pyrrolo[1',2':2,3]isoindolo[4,5,6-cd]indole-9,11(2H,7H)-dione,6a,10,11a,11b-tetrahydro-10-(1-hydroxyethylidene)-7,7-dimethyl-,(6ar,10z,11as,11br)-; 9H-Pyrrolo[1',2':2,3]isoindolo[4,5,6-cd]indol-9-one,10-acetyl-2,6,6a,7,11a,11b-hexahydro-11-hydroxy-7,7-dimethyl-,(6aR,11aS,11bR)-rel-
|
|
| CAS | 83136-88-3 | |
| PubChem CID | 135494311 | |
| ChEMBL ID | CHEMBL468766 |
*Note: the IUPAC Name was collected from PubChem.
Chemical Classification: |
|
|
|---|
——————————————————————————————————————————
NPs Species Source
| Endophyte ID | Endophyte Name | Family | Genus | Taxonomy ID | GenBank ID | Closest GenBank ID | Reference | |
|---|---|---|---|---|---|---|---|---|
| Endophyte ID | Endophyte Name | Family | Genus | Taxonomy ID | GenBank ID | Closest GenBank ID | Reference |
NPs Biological Activity
| Bioactivity Name | Target ID | Target Name | Target Type | Target Organism | Target Organism ID | Potency of Bioactivity | Activity Type | Value | Unit | Endophyte ID | Endophyte Name | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Bioactivity Name | Target ID | Target Name | Target Type | Target Organism | Target Organism ID | Potency of Bioactivity | Activity Type | Value | Unit | Endophyte ID | Endophyte Name |
NPs Physi-Chem Properties
| Molecular Weight: | 336.4 | ALogp: | 2.9 |
| HBD: | 2 | HBA: | 3 |
| Rotatable Bonds: | 0 | Lipinski's rule of five: | Accepted |
| Polar Surface Area: | 73.4 | Aromatic Rings: | 5 |
| Heavy Atoms: | 25 | QED Weighted: | 0.438 |
——————————————————————————————————————————
NPs ADMET Properties*
ADMET: Absorption
| Caco-2 Permeability: | -4.937 | MDCK Permeability: | 0.00000968 |
| Pgp-inhibitor: | 0.037 | Pgp-substrate: | 0.025 |
| Human Intestinal Absorption (HIA): | 0.018 | 20% Bioavailability (F20%): | 0.073 |
| 30% Bioavailability (F30%): | 0.407 |
——————————————————————————————————————————
ADMET: Distribution
| Blood-Brain-Barrier Penetration (BBB): | 0.03 | Plasma Protein Binding (PPB): | 99.76% |
| Volume Distribution (VD): | 0.226 | Fu: | 1.36% |
——————————————————————————————————————————
ADMET: Metabolism
| CYP1A2-inhibitor: | 0.825 | CYP1A2-substrate: | 0.592 |
| CYP2C19-inhibitor: | 0.828 | CYP2C19-substrate: | 0.341 |
| CYP2C9-inhibitor: | 0.92 | CYP2C9-substrate: | 0.956 |
| CYP2D6-inhibitor: | 0.779 | CYP2D6-substrate: | 0.308 |
| CYP3A4-inhibitor: | 0.624 | CYP3A4-substrate: | 0.589 |
——————————————————————————————————————————
ADMET: Excretion
| Clearance (CL): | 1.662 | Half-life (T1/2): | 0.393 |
——————————————————————————————————————————
ADMET: Toxicity
| hERG Blockers: | 0.01 | Human Hepatotoxicity (H-HT): | 0.31 |
| Drug-inuced Liver Injury (DILI): | 0.693 | AMES Toxicity: | 0.156 |
| Rat Oral Acute Toxicity: | 0.881 | Maximum Recommended Daily Dose: | 0.983 |
| Skin Sensitization: | 0.184 | Carcinogencity: | 0.308 |
| Eye Corrosion: | 0.003 | Eye Irritation: | 0.076 |
| Respiratory Toxicity: | 0.757 |
——————————————————————————————————————————
*Note: the ADMET properties was calculated by ADMETlab 2.0. Reference: PMID: 33893803.
Similar Compounds*
Compounds similar to EMNPD with top10 similarity:
| Similar NPs | Similar Drugs | ||||||
|---|---|---|---|---|---|---|---|
| NPs ID | NPs 2D Structure | Similarity Score | TTD ID | Drug 2D Structure | Similarity Score | ||
| ENC006110 |  |
0.568 | D0C1IW |  |
0.305 | ||
| ENC006111 |  |
0.517 | D04JCN |  |
0.304 | ||
| ENC006109 |  |
0.414 | D0X7KB |  |
0.296 | ||
| ENC001635 |  |
0.379 | D05AHE |  |
0.296 | ||
| ENC001993 |  |
0.374 | D0V3ZA |  |
0.289 | ||
| ENC002408 |  |
0.367 | D02IQY | 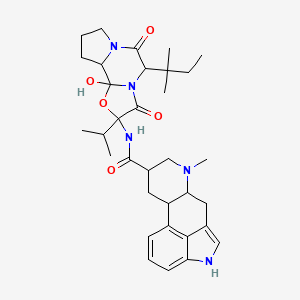 |
0.288 | ||
| ENC003722 |  |
0.307 | D09NNH |  |
0.279 | ||
| ENC003747 | 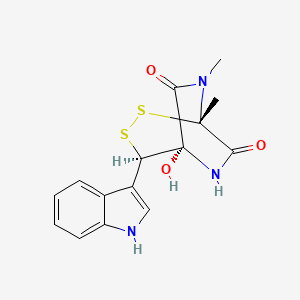 |
0.304 | D0SP3D |  |
0.278 | ||
| ENC005121 |  |
0.292 | D04EGX |  |
0.276 | ||
| ENC000091 |  |
0.289 | D01TSI |  |
0.272 | ||