NPs Basic Information
|
Name |
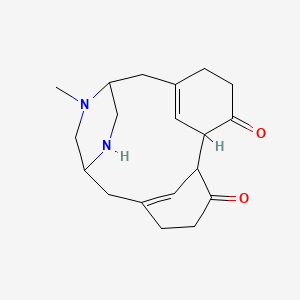
Herquline B
|
| Molecular Formula | C19H26N2O2 | |
| IUPAC Name* |
15-methyl-15,17-diazatetracyclo[12.2.2.13,7.18,12]icosa-3(20),12(19)-diene-6,9-dione
|
|
| SMILES |
CN1CC2CC3=CC(C4C=C(CCC4=O)CC1CN2)C(=O)CC3
|
|
| InChI |
InChI=1S/C19H26N2O2/c1-21-11-14-6-12-2-4-18(22)16(8-12)17-9-13(3-5-19(17)23)7-15(21)10-20-14/h8-9,14-17,20H,2-7,10-11H2,1H3
|
|
| InChIKey |
IKEHAWAEPHQJSM-UHFFFAOYSA-N
|
|
| Synonyms |
Herquline B; CHEBI:66012; Q27134516; 15-methyl-15,17-diazatetracyclo[12.2.2.1(3,7).1(8,12)]icosa-3(20),12(19)-diene-6,9-dione; 15-methyl-15,17-diazatetracyclo[12.2.2.13,7.18,12]icosa-3(20),12(19)-diene-6,9-dione
|
|
| CAS | NA | |
| PubChem CID | 9796904 | |
| ChEMBL ID | NA |
*Note: the IUPAC Name was collected from PubChem.
Chemical Classification: |
|
|
|---|
——————————————————————————————————————————
NPs Species Source
| Endophyte ID | Endophyte Name | Family | Genus | Taxonomy ID | GenBank ID | Closest GenBank ID | Reference | |
|---|---|---|---|---|---|---|---|---|
| Endophyte ID | Endophyte Name | Family | Genus | Taxonomy ID | GenBank ID | Closest GenBank ID | Reference |
NPs Biological Activity
| Bioactivity Name | Target ID | Target Name | Target Type | Target Organism | Target Organism ID | Potency of Bioactivity | Activity Type | Value | Unit | Endophyte ID | Endophyte Name | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Bioactivity Name | Target ID | Target Name | Target Type | Target Organism | Target Organism ID | Potency of Bioactivity | Activity Type | Value | Unit | Endophyte ID | Endophyte Name |
NPs Physi-Chem Properties
| Molecular Weight: | 314.4 | ALogp: | 0.3 |
| HBD: | 1 | HBA: | 4 |
| Rotatable Bonds: | 0 | Lipinski's rule of five: | Accepted |
| Polar Surface Area: | 49.4 | Aromatic Rings: | 5 |
| Heavy Atoms: | 23 | QED Weighted: | 0.698 |
——————————————————————————————————————————
NPs ADMET Properties*
ADMET: Absorption
| Caco-2 Permeability: | -4.706 | MDCK Permeability: | 0.00000205 |
| Pgp-inhibitor: | 0.001 | Pgp-substrate: | 0.012 |
| Human Intestinal Absorption (HIA): | 0.016 | 20% Bioavailability (F20%): | 0.264 |
| 30% Bioavailability (F30%): | 0.102 |
——————————————————————————————————————————
ADMET: Distribution
| Blood-Brain-Barrier Penetration (BBB): | 0.963 | Plasma Protein Binding (PPB): | 35.12% |
| Volume Distribution (VD): | 3.704 | Fu: | 65.72% |
——————————————————————————————————————————
ADMET: Metabolism
| CYP1A2-inhibitor: | 0.013 | CYP1A2-substrate: | 0.271 |
| CYP2C19-inhibitor: | 0.045 | CYP2C19-substrate: | 0.925 |
| CYP2C9-inhibitor: | 0.009 | CYP2C9-substrate: | 0.289 |
| CYP2D6-inhibitor: | 0.357 | CYP2D6-substrate: | 0.92 |
| CYP3A4-inhibitor: | 0.168 | CYP3A4-substrate: | 0.766 |
——————————————————————————————————————————
ADMET: Excretion
| Clearance (CL): | 20.573 | Half-life (T1/2): | 0.807 |
——————————————————————————————————————————
ADMET: Toxicity
| hERG Blockers: | 0.47 | Human Hepatotoxicity (H-HT): | 0.468 |
| Drug-inuced Liver Injury (DILI): | 0.046 | AMES Toxicity: | 0.13 |
| Rat Oral Acute Toxicity: | 0.206 | Maximum Recommended Daily Dose: | 0.947 |
| Skin Sensitization: | 0.4 | Carcinogencity: | 0.187 |
| Eye Corrosion: | 0.003 | Eye Irritation: | 0.007 |
| Respiratory Toxicity: | 0.77 |
——————————————————————————————————————————
*Note: the ADMET properties was calculated by ADMETlab 2.0. Reference: PMID: 33893803.
Similar Compounds*
Compounds similar to EMNPD with top10 similarity:
| Similar NPs | Similar Drugs | ||||||
|---|---|---|---|---|---|---|---|
| NPs ID | NPs 2D Structure | Similarity Score | TTD ID | Drug 2D Structure | Similarity Score | ||
| ENC004462 |  |
0.231 | D04JHN | 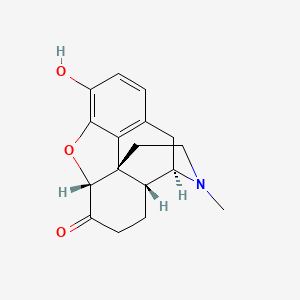 |
0.223 | ||
| ENC002636 | 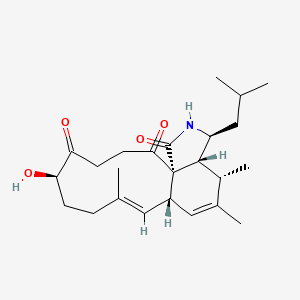 |
0.231 | D0X5KF | 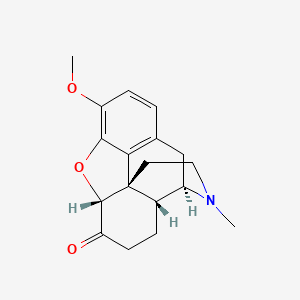 |
0.217 | ||
| ENC005810 | 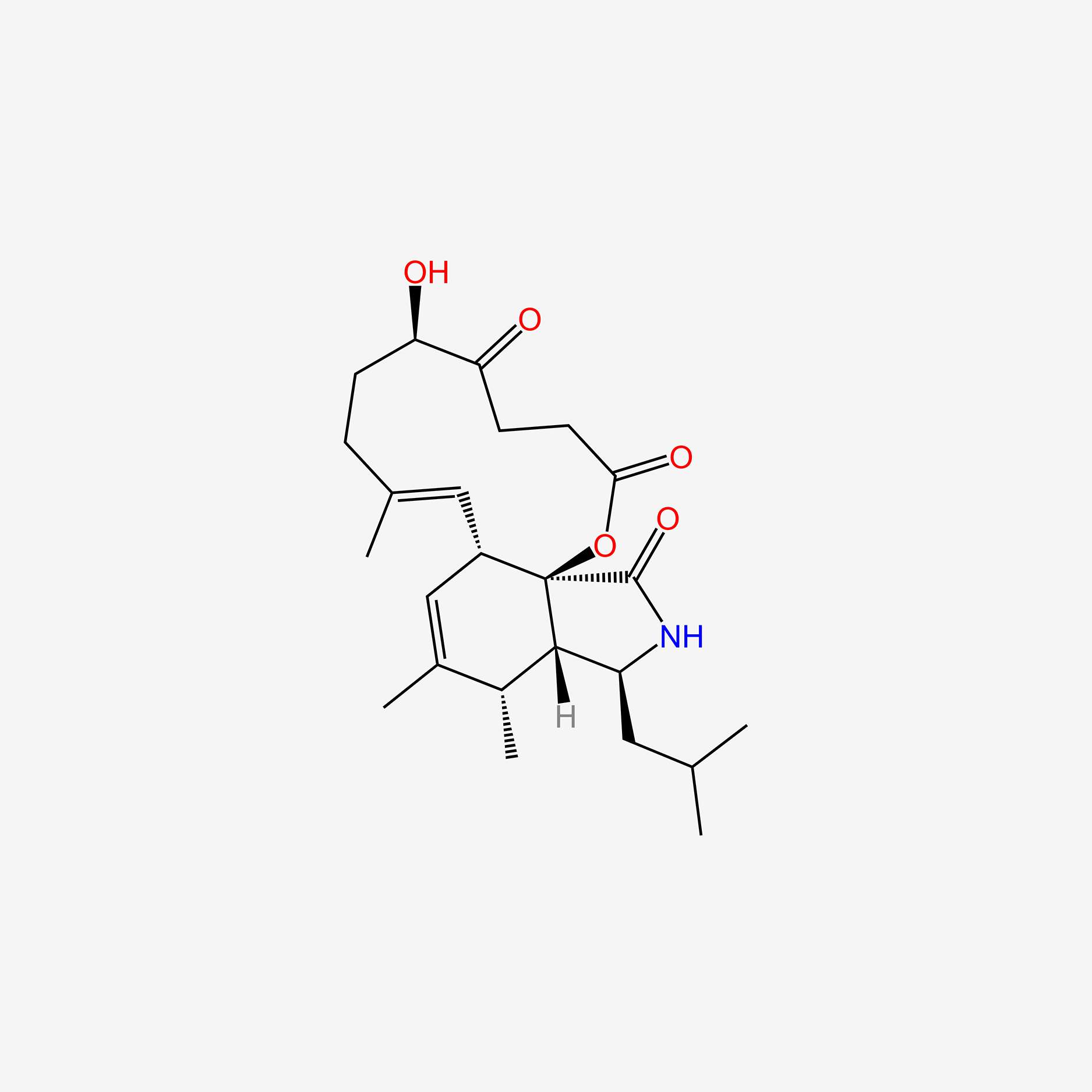 |
0.225 | D0G8BV |  |
0.215 | ||
| ENC005256 |  |
0.208 | D0F2AK |  |
0.209 | ||
| ENC003871 | 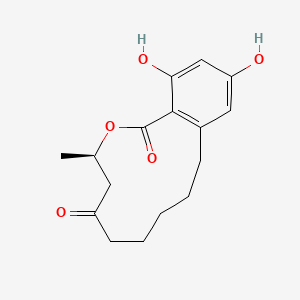 |
0.204 | D02NSF | 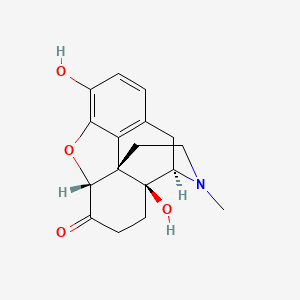 |
0.208 | ||
| ENC005002 | 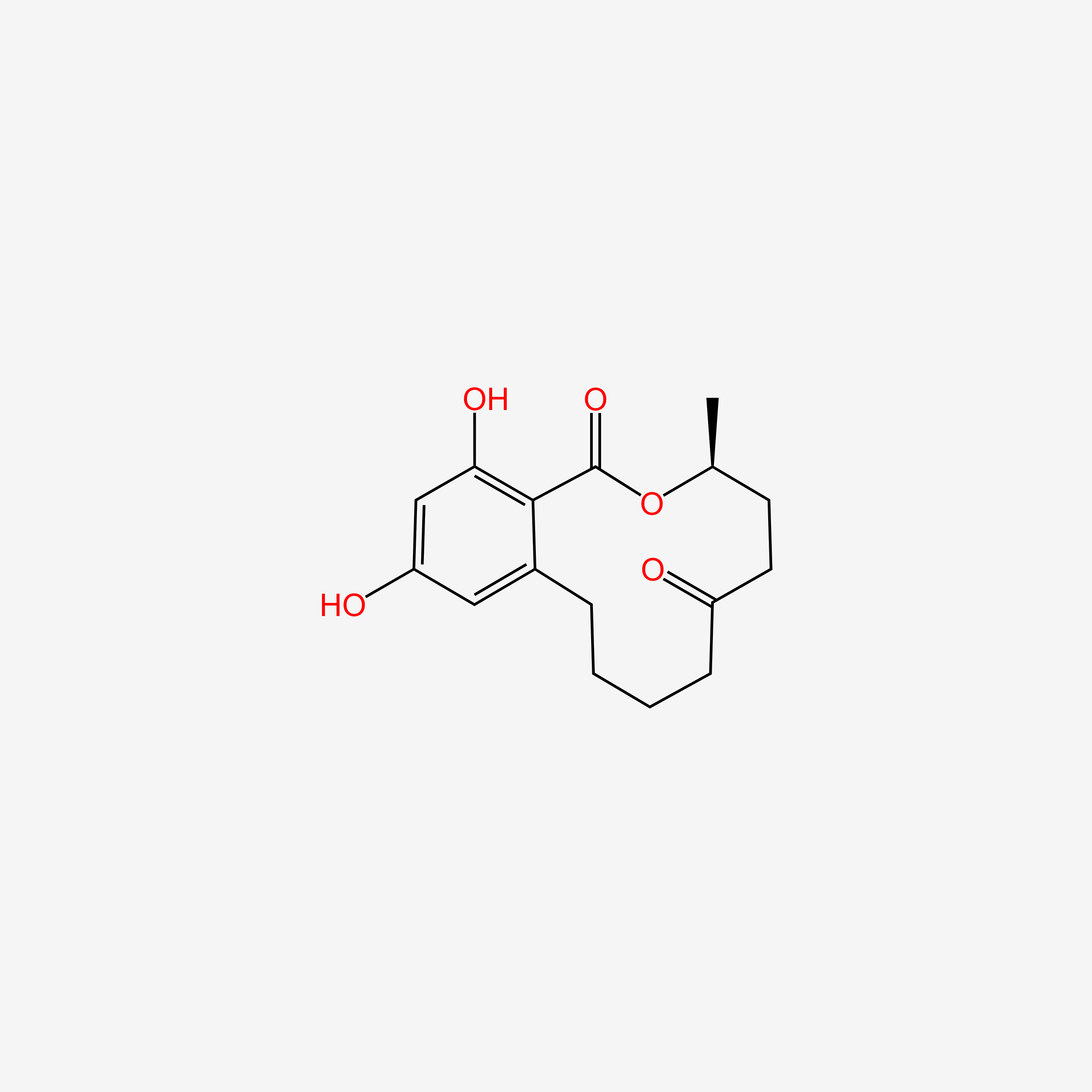 |
0.204 | D04ATM |  |
0.205 | ||
| ENC003872 | 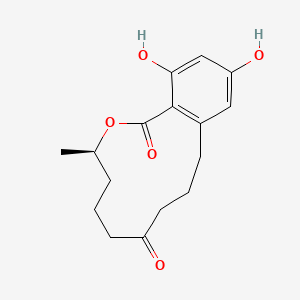 |
0.204 | D0ZV0Z | 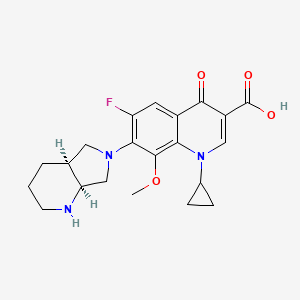 |
0.205 | ||
| ENC003673 | 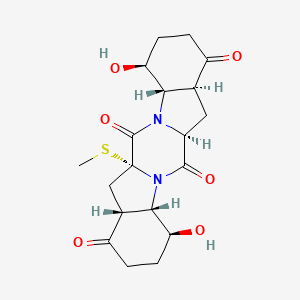 |
0.202 | D03SKD | 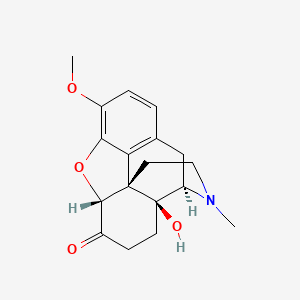 |
0.202 | ||
| ENC003715 | 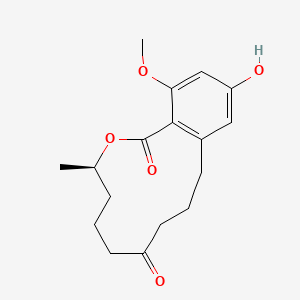 |
0.198 | D04CBI | 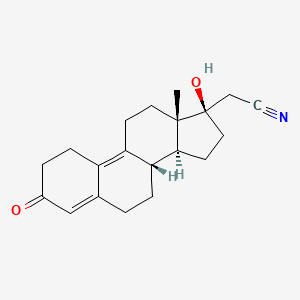 |
0.202 | ||
| ENC005001 | 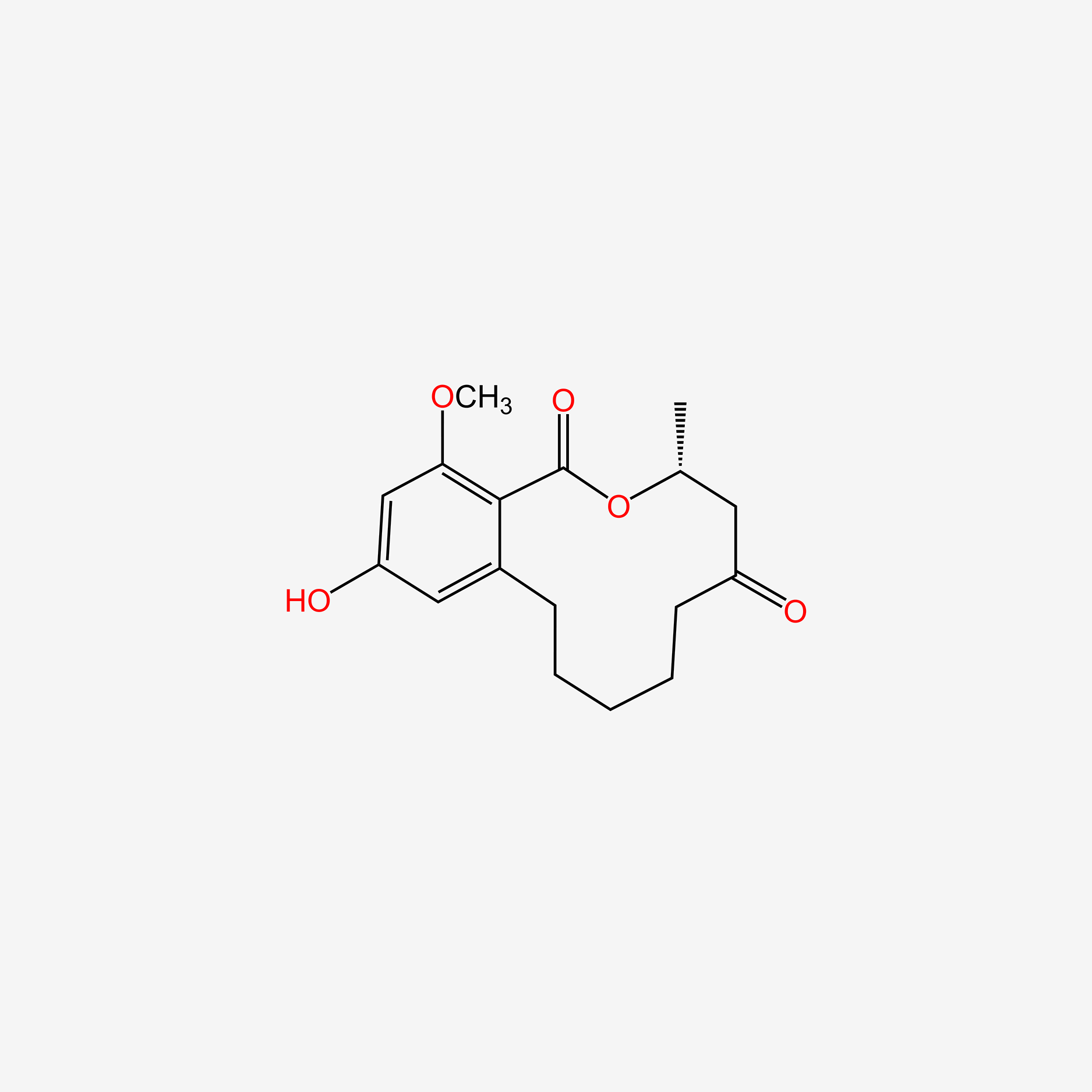 |
0.198 | D04FVU | 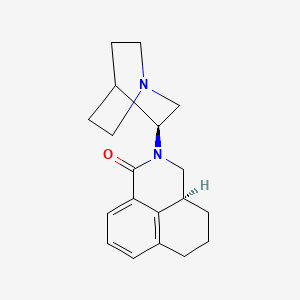 |
0.202 | ||