NPs Basic Information
|
Name |
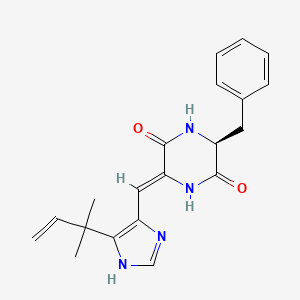
Phenylahistin
|
| Molecular Formula | C20H22N4O2 | |
| IUPAC Name* |
(3S,6Z)-3-benzyl-6-[[5-(2-methylbut-3-en-2-yl)-1H-imidazol-4-yl]methylidene]piperazine-2,5-dione
|
|
| SMILES |
CC(C)(C=C)C1=C(N=CN1)/C=C\2/C(=O)N[C@H](C(=O)N2)CC3=CC=CC=C3
|
|
| InChI |
InChI=1S/C20H22N4O2/c1-4-20(2,3)17-14(21-12-22-17)11-16-19(26)23-15(18(25)24-16)10-13-8-6-5-7-9-13/h4-9,11-12,15H,1,10H2,2-3H3,(H,21,22)(H,23,26)(H,24,25)/b16-11-/t15-/m0/s1
|
|
| InChIKey |
GWMHBVLPNWHWGW-CNYBTUBUSA-N
|
|
| Synonyms |
Phenylahistin; (-)-Phenylahistin; 200815-37-8; SCHEMBL81222; CHEMBL319291; DTXSID801336373; Q15425275; (3S,6Z)-3-benzyl-6-[[5-(2-methylbut-3-en-2-yl)-1H-imidazol-4-yl]methylidene]piperazine-2,5-dione
|
|
| CAS | 200815-37-8 | |
| PubChem CID | 9798496 | |
| ChEMBL ID | CHEMBL319291 |
*Note: the IUPAC Name was collected from PubChem.
Chemical Classification: |
|
|
|---|
——————————————————————————————————————————
NPs Species Source
| Endophyte ID | Endophyte Name | Family | Genus | Taxonomy ID | GenBank ID | Closest GenBank ID | Reference | |
|---|---|---|---|---|---|---|---|---|
| Endophyte ID | Endophyte Name | Family | Genus | Taxonomy ID | GenBank ID | Closest GenBank ID | Reference |
NPs Biological Activity
| Bioactivity Name | Target ID | Target Name | Target Type | Target Organism | Target Organism ID | Potency of Bioactivity | Activity Type | Value | Unit | Endophyte ID | Endophyte Name | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Bioactivity Name | Target ID | Target Name | Target Type | Target Organism | Target Organism ID | Potency of Bioactivity | Activity Type | Value | Unit | Endophyte ID | Endophyte Name |
NPs Physi-Chem Properties
| Molecular Weight: | 350.4 | ALogp: | 3.0 |
| HBD: | 3 | HBA: | 3 |
| Rotatable Bonds: | 5 | Lipinski's rule of five: | Accepted |
| Polar Surface Area: | 86.9 | Aromatic Rings: | 3 |
| Heavy Atoms: | 26 | QED Weighted: | 0.572 |
——————————————————————————————————————————
NPs ADMET Properties*
ADMET: Absorption
| Caco-2 Permeability: | -5.275 | MDCK Permeability: | 0.00001740 |
| Pgp-inhibitor: | 0.708 | Pgp-substrate: | 0.007 |
| Human Intestinal Absorption (HIA): | 0.006 | 20% Bioavailability (F20%): | 0.285 |
| 30% Bioavailability (F30%): | 0.003 |
——————————————————————————————————————————
ADMET: Distribution
| Blood-Brain-Barrier Penetration (BBB): | 0.022 | Plasma Protein Binding (PPB): | 98.51% |
| Volume Distribution (VD): | 0.501 | Fu: | 1.30% |
——————————————————————————————————————————
ADMET: Metabolism
| CYP1A2-inhibitor: | 0.062 | CYP1A2-substrate: | 0.218 |
| CYP2C19-inhibitor: | 0.466 | CYP2C19-substrate: | 0.062 |
| CYP2C9-inhibitor: | 0.827 | CYP2C9-substrate: | 0.967 |
| CYP2D6-inhibitor: | 0.796 | CYP2D6-substrate: | 0.07 |
| CYP3A4-inhibitor: | 0.889 | CYP3A4-substrate: | 0.859 |
——————————————————————————————————————————
ADMET: Excretion
| Clearance (CL): | 4.786 | Half-life (T1/2): | 0.922 |
——————————————————————————————————————————
ADMET: Toxicity
| hERG Blockers: | 0.039 | Human Hepatotoxicity (H-HT): | 0.292 |
| Drug-inuced Liver Injury (DILI): | 0.978 | AMES Toxicity: | 0.016 |
| Rat Oral Acute Toxicity: | 0.672 | Maximum Recommended Daily Dose: | 0.018 |
| Skin Sensitization: | 0.275 | Carcinogencity: | 0.04 |
| Eye Corrosion: | 0.003 | Eye Irritation: | 0.025 |
| Respiratory Toxicity: | 0.98 |
——————————————————————————————————————————
*Note: the ADMET properties was calculated by ADMETlab 2.0. Reference: PMID: 33893803.
Similar Compounds*
Compounds similar to EMNPD with top10 similarity:
| Similar NPs | Similar Drugs | ||||||
|---|---|---|---|---|---|---|---|
| NPs ID | NPs 2D Structure | Similarity Score | TTD ID | Drug 2D Structure | Similarity Score | ||
| ENC002895 |  |
0.560 | D0U0RZ |  |
0.288 | ||
| ENC004926 |  |
0.526 | D02PAH | 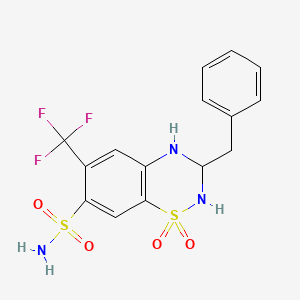 |
0.281 | ||
| ENC003988 |  |
0.524 | D01PZD |  |
0.278 | ||
| ENC004927 |  |
0.515 | D05EPM |  |
0.278 | ||
| ENC005569 |  |
0.511 | D0G1OZ | 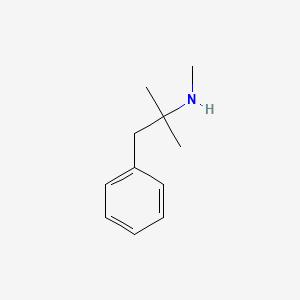 |
0.277 | ||
| ENC001957 |  |
0.511 | D0Y7RW |  |
0.277 | ||
| ENC004648 |  |
0.456 | D08UMH |  |
0.271 | ||
| ENC002631 |  |
0.448 | D02WUC | 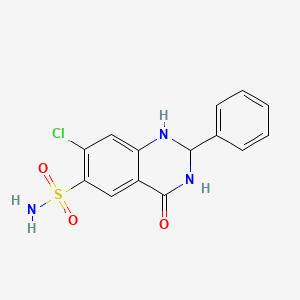 |
0.264 | ||
| ENC002717 |  |
0.424 | D09LDR | 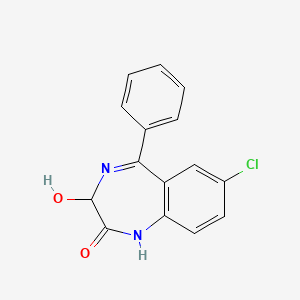 |
0.262 | ||
| ENC001910 |  |
0.415 | D0P3JU | 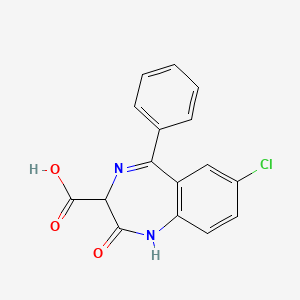 |
0.262 | ||